Manually computing spline predictions from a GAMLSS model
My goal is write down some reusable code for how to get spline predictions for unobserved x values by hand from a GAMLSS model.
Problem
Here are 98 random points from a subrange of the intelligibility dataset.
library(tidyverse)
#> Warning: package 'tibble' was built under R version 4.5.2
#> Warning: package 'tidyr' was built under R version 4.5.2
#> Warning: package 'readr' was built under R version 4.5.2
#> Warning: package 'purrr' was built under R version 4.5.2
#> Warning: package 'dplyr' was built under R version 4.5.2
#> Warning: package 'stringr' was built under R version 4.5.2
#> Warning: package 'lubridate' was built under R version 4.5.2
#> ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
#> ✔ dplyr 1.2.0 ✔ readr 2.2.0
#> ✔ forcats 1.0.1 ✔ stringr 1.6.0
#> ✔ ggplot2 4.0.2 ✔ tibble 3.3.1
#> ✔ lubridate 1.9.5 ✔ tidyr 1.3.2
#> ✔ purrr 1.2.1
#> ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
#> ✖ dplyr::filter() masks stats::filter()
#> ✖ dplyr::lag() masks stats::lag()
#> ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
suppressMessages(library(gamlss))
#> Warning: package 'nlme' was built under R version 4.5.3
select <- dplyr::select
data_intel <- tibble::tribble(
~id, ~age_months, ~mean_intelligibility,
1L, 44, 0.74,
2L, 49, 0.779,
3L, 56, 0.686,
4L, 51, 0.675,
5L, 42, 0.785,
6L, 42, 0.708,
7L, 38, 0.557,
8L, 44, 0.756,
9L, 56, 0.64,
10L, 50, 0.767,
11L, 48, 0.858,
12L, 53, 0.94,
13L, 49, 0.673,
14L, 56, 0.928,
15L, 54, 0.827,
16L, 51, 0.868,
17L, 53, 0.784,
18L, 50, 0.908,
19L, 37, 0.458,
20L, 37, 0.184,
21L, 51, 0.688,
22L, 53, 0.753,
23L, 43, 0.802,
24L, 47, 0.841,
25L, 45, 0.523,
26L, 37, 0.288,
27L, 42, 0.672,
28L, 50, 0.777,
29L, 54, 0.749,
30L, 53, 0.901,
31L, 55, 0.806,
32L, 45, 0.646,
33L, 45, 0.689,
34L, 45, 0.806,
35L, 46, 0.859,
36L, 36, 0.557,
37L, 38, 0.606,
38L, 50, 0.804,
39L, 44, 0.853,
40L, 51, 0.747,
41L, 37, 0.659,
42L, 48, 0.883,
43L, 45, 0.798,
44L, 39, 0.483,
45L, 59, 0.976,
46L, 58, 0.786,
48L, 48, 0.775,
49L, 47, 0.896,
50L, 38, 0.921,
51L, 41, 0.686,
53L, 54, 0.72,
54L, 51, 0.965,
55L, 44, 0.777,
56L, 42, 0.53,
57L, 36, 0.707,
58L, 60, 0.908,
59L, 37, 0.566,
60L, 51, 0.598,
61L, 58, 0.938,
62L, 54, 0.925,
63L, 47, 0.757,
64L, 42, 0.63,
65L, 45, 0.764,
66L, 59, 0.85,
67L, 47, 0.786,
68L, 38, 0.781,
69L, 44, 0.929,
70L, 36, 0.356,
71L, 55, 0.802,
72L, 54, 0.864,
73L, 43, 0.864,
74L, 49, 0.848,
75L, 38, 0.574,
76L, 53, 0.916,
77L, 41, 0.85,
78L, 59, 0.851,
79L, 46, 0.444,
80L, 55, 0.778,
81L, 54, 0.931,
82L, 50, 0.32,
83L, 58, 0.855,
84L, 39, 0.64,
85L, 50, 0.727,
86L, 50, 0.857,
87L, 39, 0.731,
88L, 46, 0.624,
89L, 53, 0.951,
90L, 46, 0.873,
91L, 47, 0.897,
92L, 46, 0.748,
93L, 59, 0.936,
94L, 50, 0.881,
95L, 43, 0.815,
96L, 44, 0.768,
97L, 43, 0.789,
98L, 50, 0.728,
99L, 38, 0.794,
100L, 44, 0.509
)
data_intel <- data_intel %>%
arrange(age_months)
But note that we are missing data from three ages (x values).
ggplot(data_intel) +
aes(x = age_months, y = mean_intelligibility) +
geom_point() +
geom_vline(xintercept = c(40, 52, 57), linetype = "dashed") +
scale_x_continuous(breaks = c(36, 48, 60))
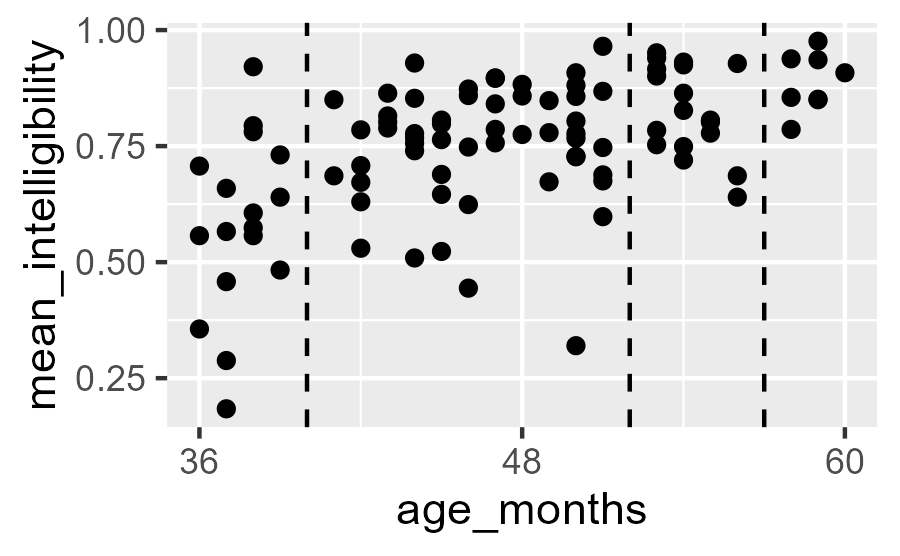
Intelligibility data but there are no observations at 40, 52, and 57 months
Here is how we would fit a beta-regression model as we did in the growth curve paper.
fit_beta_gamlss <- function(data, mu_df = 3, sigma_df = 2) {
BE <- gamlss.dist::BE
gamlss.control <- gamlss::gamlss.control
ns <- splines::ns
model <- wisclabmisc::mem_gamlss(
mean_intelligibility ~ ns(age_months, df = mu_df),
sigma.formula = ~ ns(age_months, df = sigma_df),
family = BE(),
data = data,
control = gamlss.control(trace = FALSE)
)
model$.user$mu_df <- mu_df
model$.user$sigma_df <- sigma_df
model$.user$mu_basis <- ns(data$age_months, mu_df)
model$.user$sigma_basis <- ns(data$age_months, sigma_df)
model
}
model <- fit_beta_gamlss(data_intel, mu_df = 3, sigma_df = 2)
Then we can get centiles pretty easily for the existing x values. But note the three gaps on the percentile line.
centiles <- data_intel %>%
select(age_months) %>%
wisclabmisc::predict_centiles(model)
ggplot(data_intel) +
aes(x = age_months, y = mean_intelligibility) +
geom_point() +
geom_point(
aes(y = c10),
data = centiles,
color = "skyblue3",
size = 10,
shape = "-"
) +
scale_x_continuous(breaks = c(36, 48, 60))
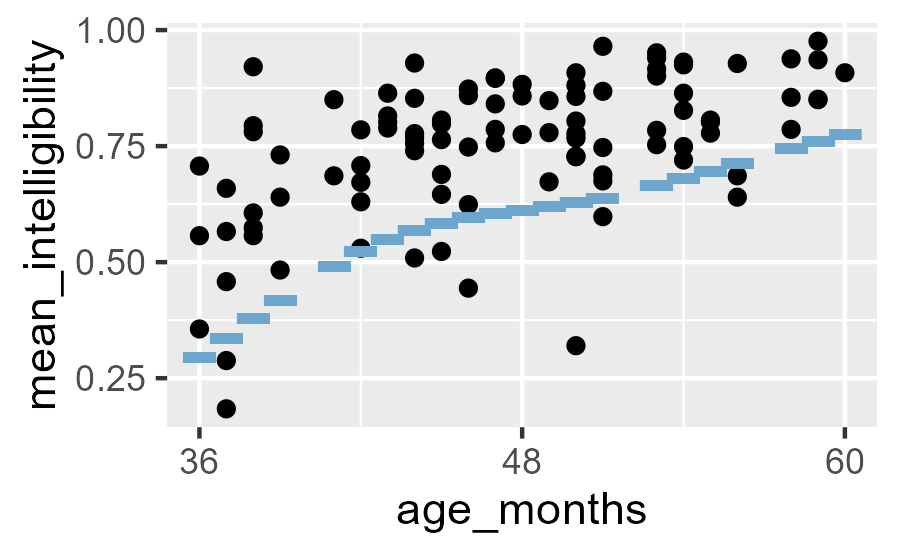
Intelligibility data with points from the 10th percentile line
If we ask GAMLSS to give us centiles for those gaps—herein lies the problem—we might get a warning message like the following:
centiles2 <- data_intel %>%
select(age_months) %>%
tidyr::expand(age_months = tidyr::full_seq(age_months, 1)) %>%
wisclabmisc::predict_centiles(model)
#> Warning in predict.gamlss(obj, what = "mu", newdata = newx, type = "response", : There is a discrepancy between the original and the re-fit
#> used to achieve 'safe' predictions
#>
I don’t want to mess around in unsafe territory with GAMLSS. (What if they change how they handle this problem in the future?) So I would like to estimate values for those missing x values from the spline by hand.
Solution
Use predict() on the spline object to get the values
of the basis functions at the new x values. I stashed
the original spline object inside of the model.
mu_basis <- model$.user$mu_basis
ages_to_predict <- data_intel %>%
tidyr::expand(age_months = tidyr::full_seq(age_months, 1))
mu_basis2 <- predict(mu_basis, newx = ages_to_predict$age_months)
library(patchwork)
tjmisc::ggmatplot(mu_basis) +
tjmisc::ggmatplot(mu_basis2)
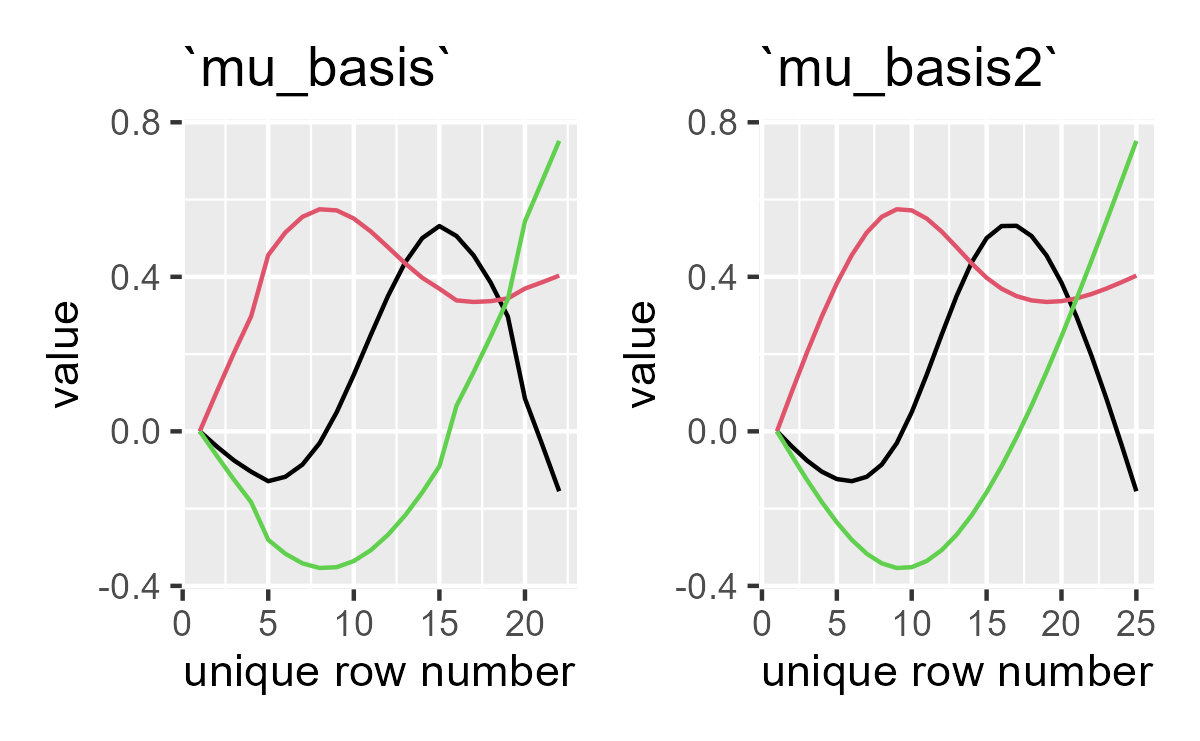
Basis functions for observed data (left) and the age range (right). The full age range is a little bit smoother than the observed data.
We do the following to get model parameters at each age. This code is the point of this note.
mu_basis <- model$.user$mu_basis
sigma_basis <- model$.user$sigma_basis
newx <- ages_to_predict$age_months
mu_basis2 <- cbind(1, predict(mu_basis, newx = newx))
sigma_basis2 <- cbind(1, predict(sigma_basis, newx = newx))
manual_centile <- tibble::tibble(
age_months = newx,
# multiply coefficients and undo link function
mu = plogis(mu_basis2 %*% coef(model, what = "mu"))[, 1],
sigma = plogis(sigma_basis2 %*% coef(model, what = "sigma"))[, 1],
# now take centiles
c10 = gamlss.dist::qBE(.1, mu, sigma)
)
manual_centile
#> # A tibble: 25 × 4
#> age_months mu sigma c10
#> <dbl> <dbl> <dbl> <dbl>
#> 1 36 0.522 0.341 0.294
#> 2 37 0.560 0.334 0.336
#> 3 38 0.596 0.328 0.378
#> 4 39 0.630 0.321 0.418
#> 5 40 0.661 0.315 0.456
#> 6 41 0.688 0.309 0.491
#> 7 42 0.712 0.304 0.522
#> 8 43 0.731 0.298 0.548
#> 9 44 0.745 0.293 0.569
#> 10 45 0.755 0.288 0.584
#> # ℹ 15 more rows
Let’s look at our manually computed 10th percentile versus the package-provided one.
ggplot(data_intel) +
aes(x = age_months, y = mean_intelligibility) +
geom_point() +
geom_point(
aes(y = c10),
data = manual_centile,
color = "orange",
size = 10,
shape = "+"
) +
geom_point(
aes(y = c10),
data = centiles,
color = "skyblue3",
size = 10,
shape = "-"
) +
scale_x_continuous(breaks = c(36, 48, 60))
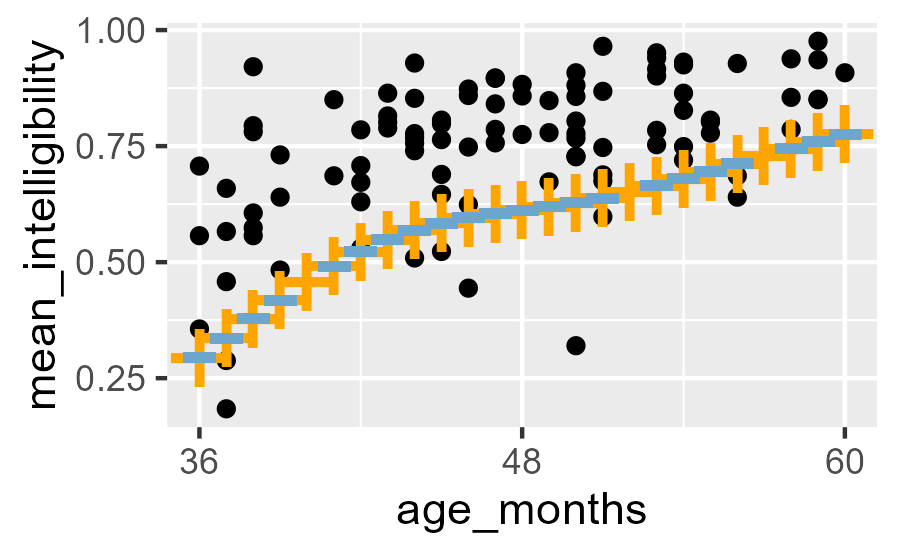
Intelligibility data with points from the 10th percentile line provided by `centiles.pred()` (blue hyphens) and computed by hand (orange pluses).
Leave a comment