Confidence interval overlap and p-values
Two 95% confidence intervals can overlap and still have a statistically significant difference at p = .05. I want to refresh my memory on what p-values are implied by different degrees of overlap and simulate the rules of thumb myself.
Cumming (2008) provides a summary of some visual rules of thumb. (This is the same author of 2014 The New Statistics manifesto.) Here are the main points I wanted to recall:
- Two 95% intervals can overlap by a small amount at p = .01.
- Two 83.5% intervals will just touch at p = .05.
- Two 95% intervals can overlap up to half the length of an interval whisker at p = .05.
We simulate data sets that yield a target p-value.
simulate_group_data_for_given_p_value <- function(
p_target,
n_a = 40,
n_b = NULL,
sd_a = 1,
sd_b = NULL,
seed = NULL
) {
if (!is.null(seed)) withr::local_seed(seed)
n_b <- n_b %||% n_a
sd_b <- sd_b %||% sd_a
data_a <- rnorm(n_a, 0, sd_a)
data_b <- rnorm(n_b, 0, sd_b)
get_p_val <- function(diff) t.test(data_a, data_b + diff)[["p.value"]]
target_diff <- uniroot(
f = function(x) get_p_val(x) - p_target,
interval = c(0, 5 * sd_a),
tol = p_target / 1000,
extendInt = "yes"
)
data.frame(
group = c(rep("a", n_a), rep("b", n_b)),
value = c(data_a, data_b + target_diff$root),
target_p_value = paste0("p = ", p_target)
)
}
Test the data simulation functions.
data1 <- simulate_group_data_for_given_p_value(.05, seed = 20250723)
t.test(value ~ group, data1) |>
broom::tidy() |>
str()
#> tibble [1 × 10] (S3: tbl_df/tbl/data.frame)
#> $ estimate : num -0.443
#> $ estimate1 : num 0.026
#> $ estimate2 : num 0.469
#> $ statistic : Named num -1.99
#> ..- attr(*, "names")= chr "t"
#> $ p.value : num 0.05
#> $ parameter : Named num 77.9
#> ..- attr(*, "names")= chr "df"
#> $ conf.low : num -0.887
#> $ conf.high : num -8.44e-07
#> $ method : chr "Welch Two Sample t-test"
#> $ alternative: chr "two.sided"
data2 <- simulate_group_data_for_given_p_value(.01, seed = 20250723)
t.test(value ~ group, data2) |>
broom::tidy() |>
str()
#> tibble [1 × 10] (S3: tbl_df/tbl/data.frame)
#> $ estimate : num -0.588
#> $ estimate1 : num 0.026
#> $ estimate2 : num 0.614
#> $ statistic : Named num -2.64
#> ..- attr(*, "names")= chr "t"
#> $ p.value : num 0.01
#> $ parameter : Named num 77.9
#> ..- attr(*, "names")= chr "df"
#> $ conf.low : num -1.03
#> $ conf.high : num -0.145
#> $ method : chr "Welch Two Sample t-test"
#> $ alternative: chr "two.sided"
Create a grid of CIs and plot them. Hmisc::smean.cl.normal() is used by
ggplot2 to compute the CI, and this function “the sample mean and
lower and upper Gaussian confidence limits based on the
t-distribution”.
library(tidyverse)
#> Warning: package 'tibble' was built under R version 4.5.2
#> Warning: package 'tidyr' was built under R version 4.5.2
#> Warning: package 'readr' was built under R version 4.5.2
#> Warning: package 'purrr' was built under R version 4.5.2
#> Warning: package 'dplyr' was built under R version 4.5.2
#> Warning: package 'stringr' was built under R version 4.5.2
#> Warning: package 'lubridate' was built under R version 4.5.2
#> ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
#> ✔ dplyr 1.2.0 ✔ readr 2.2.0
#> ✔ forcats 1.0.1 ✔ stringr 1.6.0
#> ✔ ggplot2 4.0.2 ✔ tibble 3.3.1
#> ✔ lubridate 1.9.5 ✔ tidyr 1.3.2
#> ✔ purrr 1.2.1
#> ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
#> ✖ dplyr::filter() masks stats::filter()
#> ✖ dplyr::lag() masks stats::lag()
#> ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
mean_ci <- function(x, width) {
df <- ggplot2::mean_cl_normal(x, conf.int = width)
df[[".width"]] <- width
df
}
data <- bind_rows(
data1,
data2,
simulate_group_data_for_given_p_value(.005, seed = 20250723)
)
cis <- data |>
group_by(group, target_p_value) |>
reframe(
ci = list(mean_ci(value, .95), mean_ci(value, .835))
) |>
tidyr::unnest(cols = ci)
ggplot(cis) +
aes(x = factor(.width), color = group) +
geom_pointrange(
aes(ymin = ymin, y = y, ymax = ymax),
position = position_dodge(width = .25)
) +
facet_wrap(~target_p_value) +
guides(color = "none") +
scale_x_discrete(
"Confidence interval",
labels = function(x) scales::label_percent(.1)(as.numeric(x))
) +
labs(y = NULL) +
theme_grey(base_size = 14)
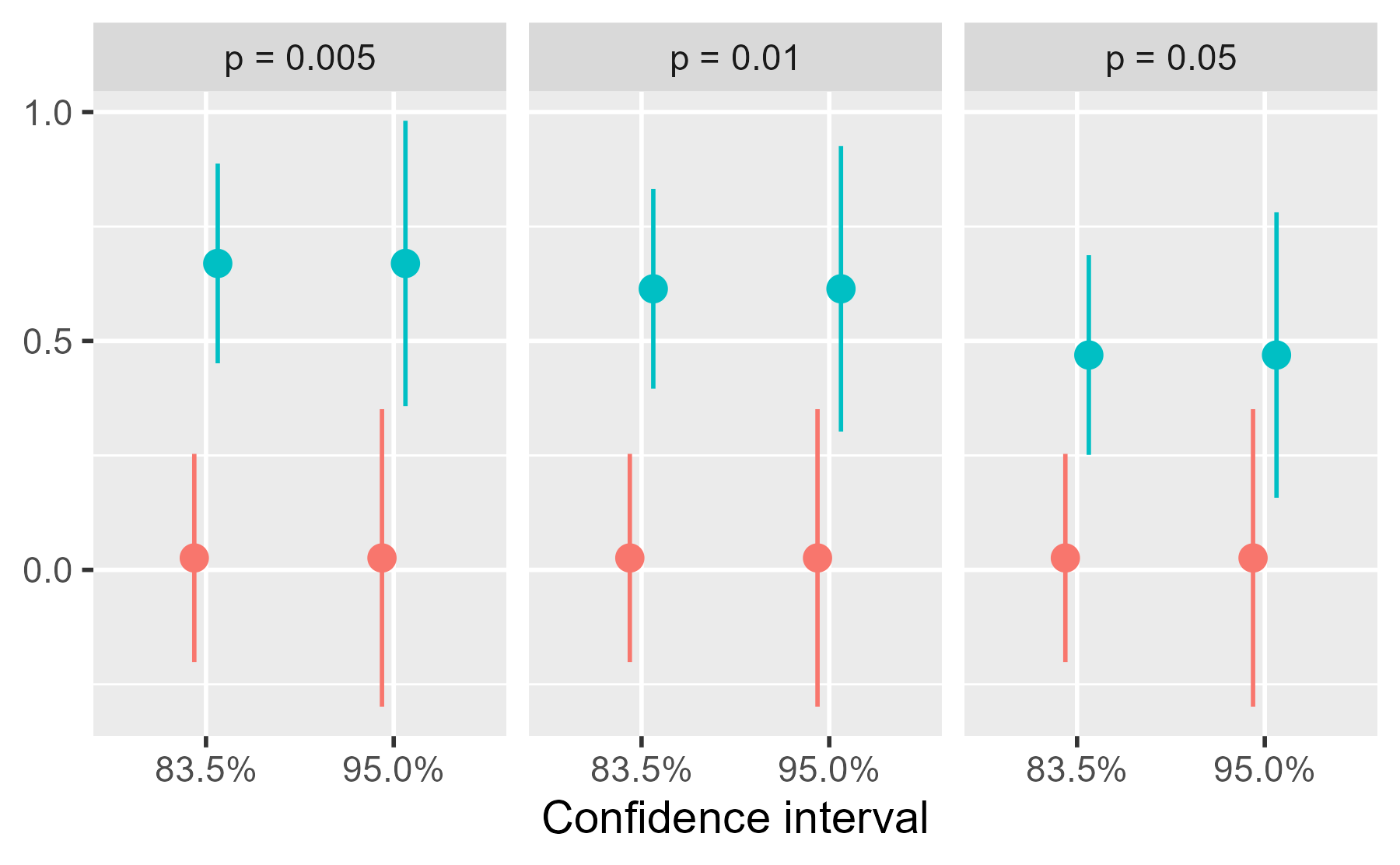
center
I was also intrigued by the following remark:
Sall [28] described an ingenious variation of overlap: Around any mean, draw a circle with radius equal to the margin of error of the 95 per cent CI. Sall showed that if two such circles overlap so that they intersect at right angles, p = 0.05 for the comparison of the two means.
So, when p = .05, the tangents at the intersections are right angles (?).
data_circles <- data1 |>
bind_rows(
simulate_group_data_for_given_p_value(.10, seed = 20250723),
simulate_group_data_for_given_p_value(.20, seed = 20250723)
) |>
group_by(group, target_p_value) |>
reframe(
ci = mean_ci(value, .95)
) |>
tidyr::unnest(cols = ci) |>
filter(.width == .95) |>
mutate(
radius = (ymax - ymin) / 2
) |>
rowwise() |>
mutate(
angle = list(seq(0, 2 * pi, length.out = 100))
) |>
ungroup() |>
tidyr::unnest(cols = angle) |>
mutate(
circle_x = cos(angle) * radius,
circle_y = y + sin(angle) * radius
)
ggplot(data_circles) +
geom_point(aes(x = 0, y = y)) +
geom_path(aes(group = y, x = circle_x, y = circle_y)) +
facet_wrap(~target_p_value) +
coord_fixed()
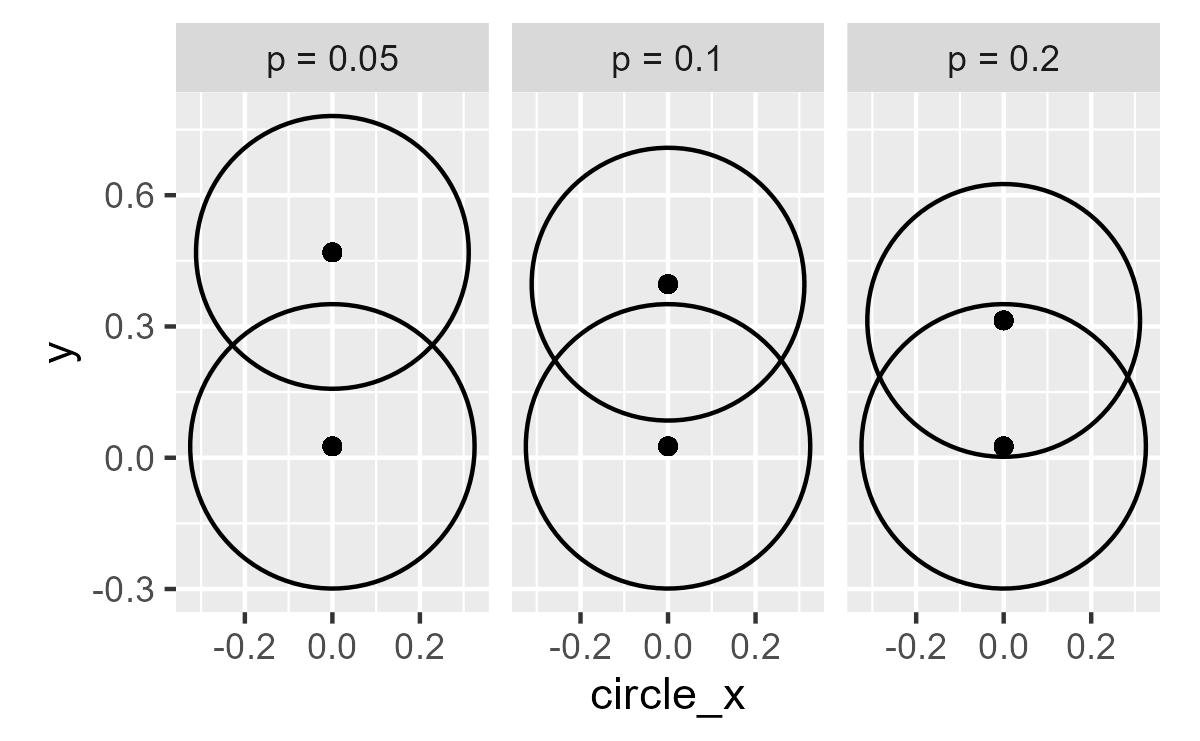
center
Leave a comment